Microbiologists have identified over a hundred phyla of bacteria, groups that are astonishingly disparate in their cell architecture, physiology, biochemistry, and most importantly, their ecology. Our lab has been interested for decades in the first step toward this ecological diversification. This is speciation, where one lineage splits into two “ecotypes” that can coexist indefinitely as a result of their ecological divergence.

Because of the very low frequency of recombination in bacteria (usually around the rate of mutation), the process of speciation appears simpler and faster in bacteria than in the highly sexual animals. There appears to be no need for either evolution of sexual isolation or geographic isolation in bacterial speciation. Indeed, microbial ecologists are finding evidence of sympatric speciation (among populations that are within easy dispersal range of one another) in nature and in laboratory evolution experiments.
We recently sought to test a novel hypothesis about the propensity for speciation in bacteria—that a lineage might split into two ecotypes that can coexist indefinitely (“cladogenesis” or speciation) faster than the lineage improves its adaptations without splitting (“anagenesis”). Our model system was a set of ten replicate communities of Bacillus subtilis growing in an extremely rich liquid medium. These communities were founded with two genetically marked strains derived from lab strain 168, with no other species present. By analyzing the pattern of frequency changes in the marked strains, we could tell whether the first adaptive mutation within a community led to a cladogenesis or an anagenesis event. Remarkably, we found that in a majority of communities there was a splitting of the original population into multiple ecologically distinct populations (or “ecotypes”) before there was even a single adaptive improvement within the original population. Thus, speciation appeared to occur at least as fast as adaptive improvement within the original ecotype. Moreover, we found that speciation was occurring in a matter of several days or less.
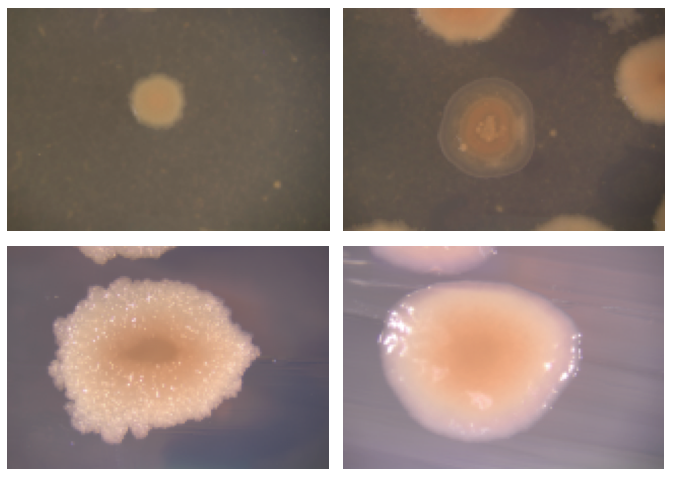
We were able to corroborate the existence of multiple ecologically distinct populations through various approaches. First, we found that in several of our communities there was one ecotype that colonized the surface while another lived in the water column (second figure). Also, we found a diversity of colony morphology types within each community we sampled (first figure), and we showed in head-to-head competition experiments that different colony types were always able to coexist in the medium. We were amazed at how well B. subtilis was poised to diversify quickly.
The absence of other species may have accelerated the rate of speciation in the microcosms. This is because other soil species may already hold the niches that B. subtilis claimed in our experiments. Our study most directly simulates evolution following colonization of sterilized soil, for example following a volcanic eruption, colonization of a Mars-like planet lacking competing organisms (if it contains organic material), or colonization of sterilized food. While the potential for speciation may be much lower in natural soils teeming with other bacteria, we have established a baseline for the potential for speciation. In the absence of other organisms, one clone can diversify into multiple ecotypes at least as fast as a single ecotype can be improved.
Further reading: